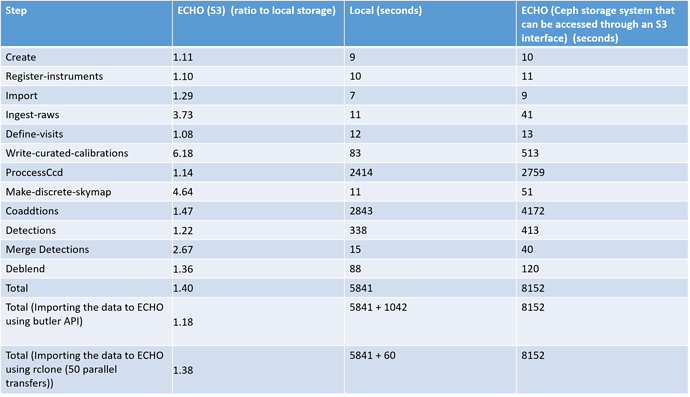
We are committed to supporting object stores for processing and welcome any feedback you give us. I will have to sort out file caching at some point because we are planning on adding quantum clustering to the workflow graph so that a single job can run multiple quanta – this will make use of caching by noting that the next quantum will be able to read the output locally from the previous quantum without having to fetch it from the object store.
I am also wondering whether we should add asyncio support to ButlerURI and allow a bulk butler.put that can parallelize the file transfers on job completion. Allowing a bulk ingest that can use ButlerURI.transfer_from() with asyncio might be useful as well.
There are two formats of index files so you get to choose. One stores the translated metadata and the other stores all the FITS headers. Translated metadata is he smallest and fastest to process but we probably will use the raw header option in the index because that form allows us to modify the header translation and reingest without rewriting the index file (so the index file can become a permanent read-only artifact).
We haven’t used these index files “in real life” and there has been talk of allowing an option that writes the index files to a parallel directory tree rather than storing them directly alongside the raw data. Doing that makes it easier to delete and rebuild them without having to touch the curated raw folders.
A lot of this also assumes that these index files are created incrementally as new data arrives (and so the headers can be read during data transfer) but of course creating these index files from pre-existing data in buckets still requires download of the entire file. I have been considering the ability to read the first N-bytes from the file to minimize the download. This should work fine for LSSTCam data but for DECam it won’t work because DECam files store multiple detectors in a single file.
Our plan for the IDF is to have a butler client/server that uses the science platform A&A system to access it and returns signed URLs that clients can use for reading from and writing to the object store. Science users will not have direct access to the registry SQL database.

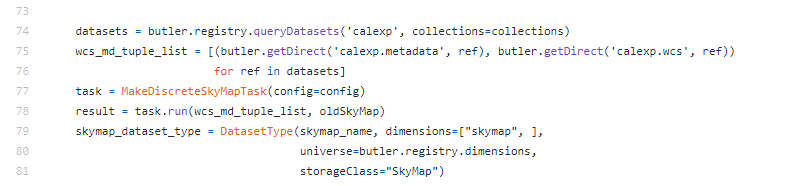
 , cheers for the correction
, cheers for the correction